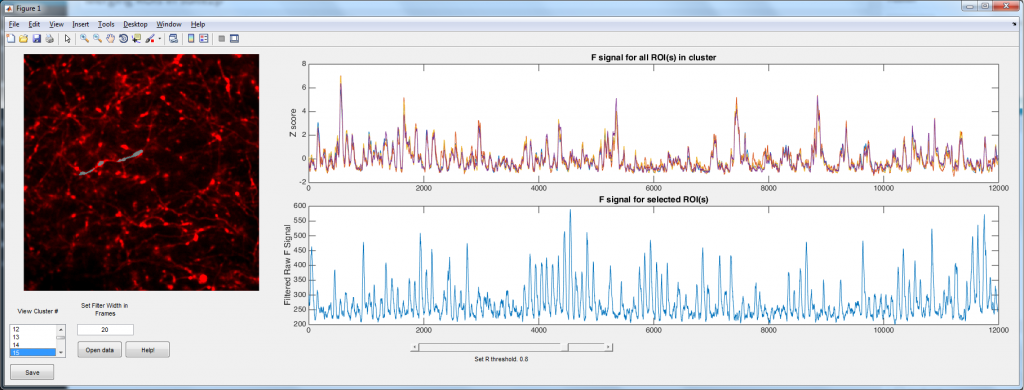
Suite2p is a wonderful Matlab toolbox written by Marius Pachitariu for analyzing population calcium imaging data. It uses a number of computational tricks to automate and accelerate the process (so no more drawing regions of interest (ROIs) by hand!). However, I spend most of my time imaging dendrites and axons, and here suite2p has a problem. Suite2p uses a heuristic that is looking for approximately elliptical ROIs, and hence it tends to split axons/dendrites into a large number smaller ROIs. The problem was simple: how can we merge the ROIs belonging to single cells? Well I used the logic that ROIs that belong to the same neuron should have highly correlated calcium signals (yes, I can imagine a situations where this wont be the case in dendrites, but bAPs will still dominate the calcium trace 99.9% of the time). Hence I simply correlate each ROI with every other ROI. ROIs with a correlation coefficient above some user settable threshold are considered to be part of the same process.
The main script is available here, and it requires distinguishable_colors.m (which in turn requires the image processing toolbox I believe).
The code is relatively well documented/commented, and there is even a ‘Help!’ button. If anyone has any problems with it, please let me know.
Hi Bill,
Great to see your post. I’m doing axonal calcium imaging so this function is exactly what I want. But do you also have python code?
Sean
Hi Sean, I wrote this back when suite2p was Mtlab only. I don’t have it in Python. And I’m not going to lie, I hate doing GUIs in Python, so I’m not about to jump into it. But the core functionality is pretty simple
1) First filter the data (Box car is fine, but do something fancy if you want)
2) calculate the correlation coefficient between each ROI and every other ROI.
3) If the coefficient is above some threshold, declare the ROIs to be the same cell. (I found R typically needed to be about >0.7-0.8 for the ROIs to be the same.)